Project Category: Chemical

About Our Project
Plastic pollution is well-recognized as a global problem. The massive amounts of plastic waste in the environment threaten ecosystems, food chains, and health. The textile industry is a major contributor to plastic waste, with 35% of the primary plastic in the ocean coming from clothing products. 60% of fabric textiles are composed of plastics, such as polyester and nylon, which are non-biodegradable and accumulate in the environment. In order to effectively alleviate the effects of plastic pollution, preventative approaches are required such as adopting alternative materials as replacements for plastic-containing products. To accomplish the goal of reducing environmental waste, biologically derived materials pose significant advantages. Structural proteins, like spider silk, are both biodegradable and mechanically versatile making them an environmentally-friendly alternative to plastic material employed in textile applications.
Although the holistic design of spider silk manufacturing processes is lacking, previous work has identified limiting process components. Our group has identified three aspects stifling design performance: the efficiency of silk protein synthesis, the economic efficiency of the silk recovery, and the production capacity of the fiber preparation process. The scope of this report evaluates prior process selections and alternative unit operations for these design aspects.
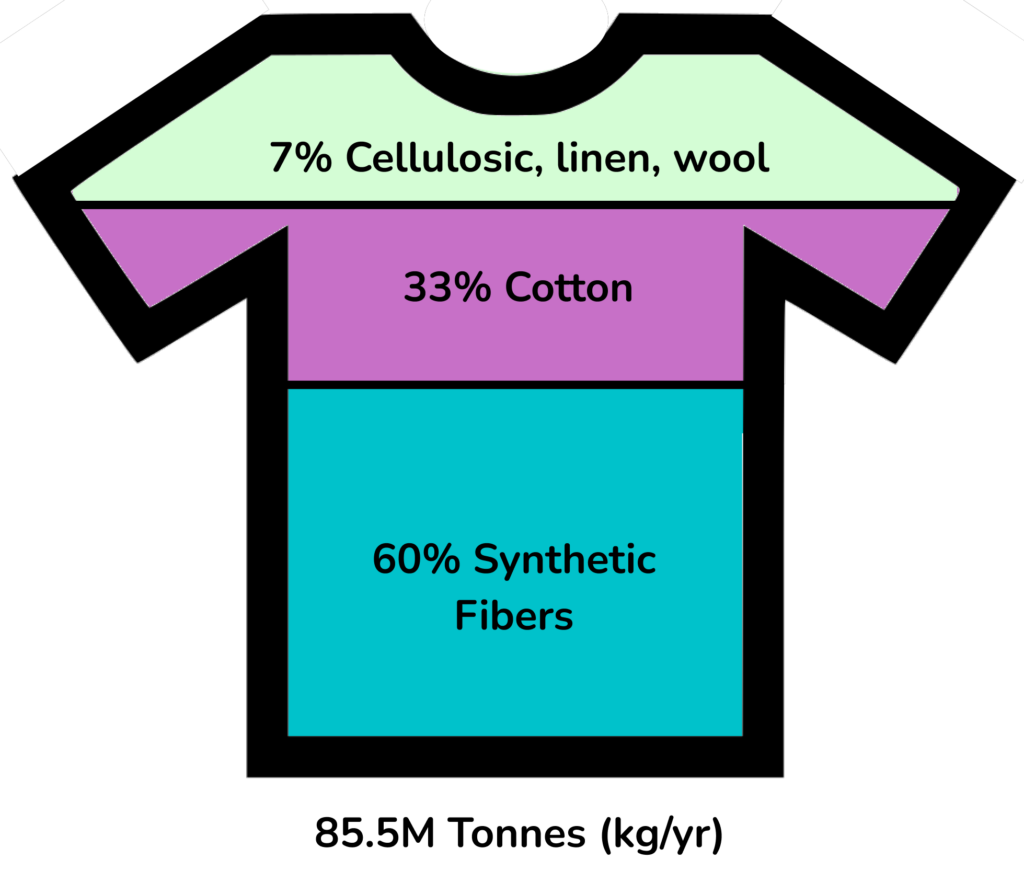
clothing based textiles for the global industry
Details About Our Design
HOW OUR DESIGN ADDRESSES PRACTICAL ISSUES
Microplastics are a major source of plastic pollution, which contributes to the vast amounts of plastic found in oceans. The accumulation of plastic in oceans has detrimental effects on seabirds and marine animals as they can easily ingest these materials or become entangled in them. Not only this, microplastics have recently been discovered in the human bloodstream. While we know that microplastics are increasingly being found in living organisms, we don’t know the long-term effects these materials may cause. By designing a biodegradable protein-based fiber, Silk Sense can help reduce the detrimental effects that microplastics may cause.
Currently, silk protein-based textiles are primarily used in luxury products and therefore, are not commonly found in everyday clothing products. Because of this, making the switch between plastic textiles and spider silk-based textiles is difficult as plastic textiles are a much cheaper alternative. Silk Sense is working to reduce the cost of this unique material, in hopes that by doing so, the material becomes more widely used.
WHAT MAKES OUR DESIGN INNOVATIVE
Silk Sense is determined to increase the accessibility of spider silk protein textiles, in hopes of reducing microplastic pollution. Although the use of spider silk in textile materials has long been of interest to many companies and researchers, many only create this product in lab settings. Few companies have worked to design an upscaled plant for the mass production of this product. While few companies have successfully created a spider silk protein-based textile, this product is not commercially available and is commonly used inexpensive luxury products. At Silk Sense, we’ve worked to create a feasible design that helps make this product one step further to being accessible on a large scale.
In terms of the product itself, our product is a high purity, strong tensile material which provides a biodegradable, renewable alternative to currently used plastic textiles. By implementing this material into clothing, Silk Sense can progressively help the world reduce its reliance on plastic materials.
WHAT MAKES OUR DESIGN SOLUTION EFFECTIVE
Although the term ‘silk’ may not sound like a strong alternative to plastic textiles, spider silk has actually shown strong mechanical properties such as elasticity and tensile strength. Additionally, these mechanical properties have shown to be stronger than those in currently used plastics such as nylon and kevlar, making silk proteins a highly effective alternative.
Along with this, Silk Sense has designed a process that creates a silk protein fiber of 87% purity. By achieving this purity, Silk Sense can ensure the strong mechanical properties mentioned above are maintained in our final material.
HOW WE VALIDATED OUR DESIGN SOLUTION
The four major units of the designed plant were individually validated using a combination of literature, engineering calculations, and simulations. For more information on the validation of each unit, see the individual unit tabs.
FEASIBILITY OF OUR DESIGN SOLUTION
Our plant design is economically feasible, which is reflected in the rate of return of 6.06% and a payout period of 14.3%. Various changes in the unit designs and material expenses were closely examined to improve the economics in the initial state of our design. The initial design of the plant required significant use of raw materials, in which alternative designs for the microbe growth and purification unit were used to reduce expenses. In the fiber spinning unit, the geometric design of the spinneret was optimized to accommodate the highest velocity it can take while retaining the shear needed to make the material durable and spin continuously. Increasing the output velocity of the spinneret enabled our plant design to raise production, which effectively increased our revenue as well.
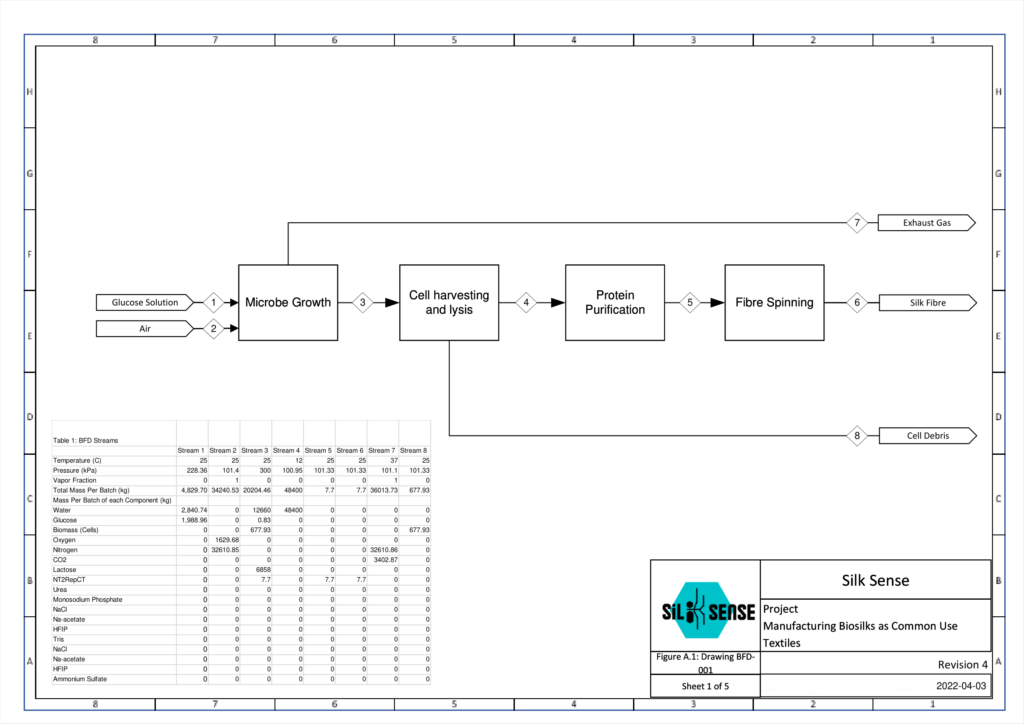
Overall Process Design
Microbe Growth and Analysis
Cell Harvesting and Lysis
Protein Recovery and Purification
Fibre Preparation
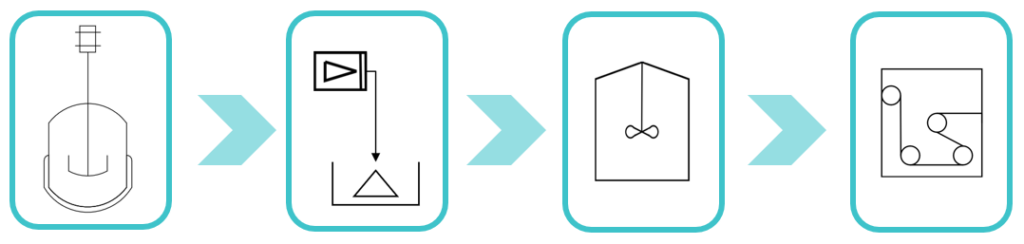
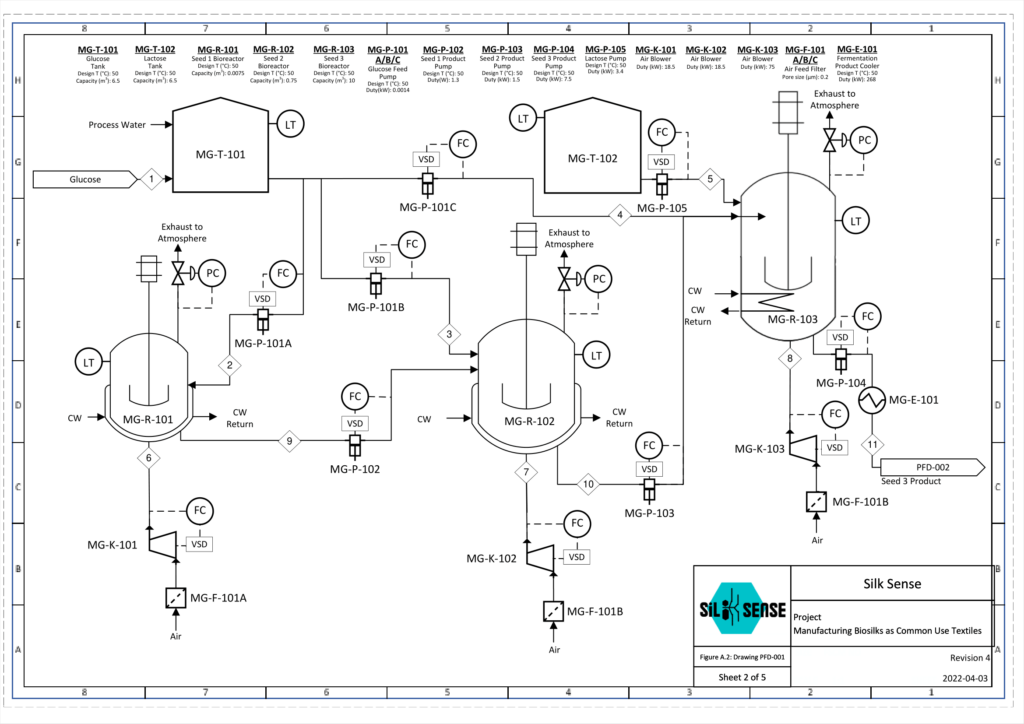
Microbe Growth and Protein Synthesis
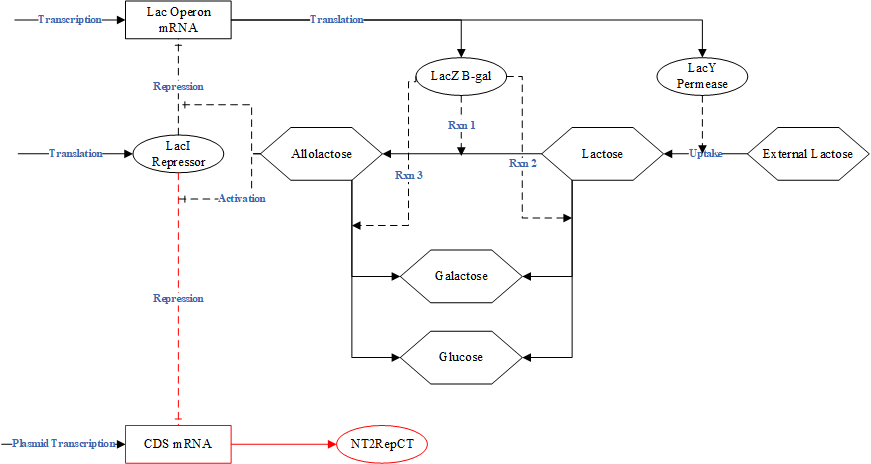
The recombinant spider silk protein, NT2RepCT, which makes up the spider silk fiber, is synthesized by genetically engineered E. coli BL21 (DE3). To activate protein synthesis, the lactose induction system is utilized.
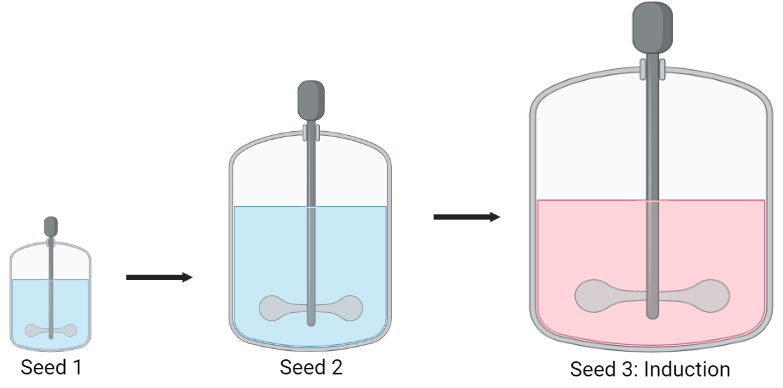
To obtain an industrially relevant quantity, the modified organism is passed through a seed train consisting of three bioreactors of increasing sizes. Seed 1 and 2 consist of 7.5 L and 750 L bioreactors to allow a small amount of modified E. coli to be substantially increased. In the final 10000 L bioreactor, lactose is added as a chemical inducer which initiates synthesis of the silk protein inside the cells.
Validation
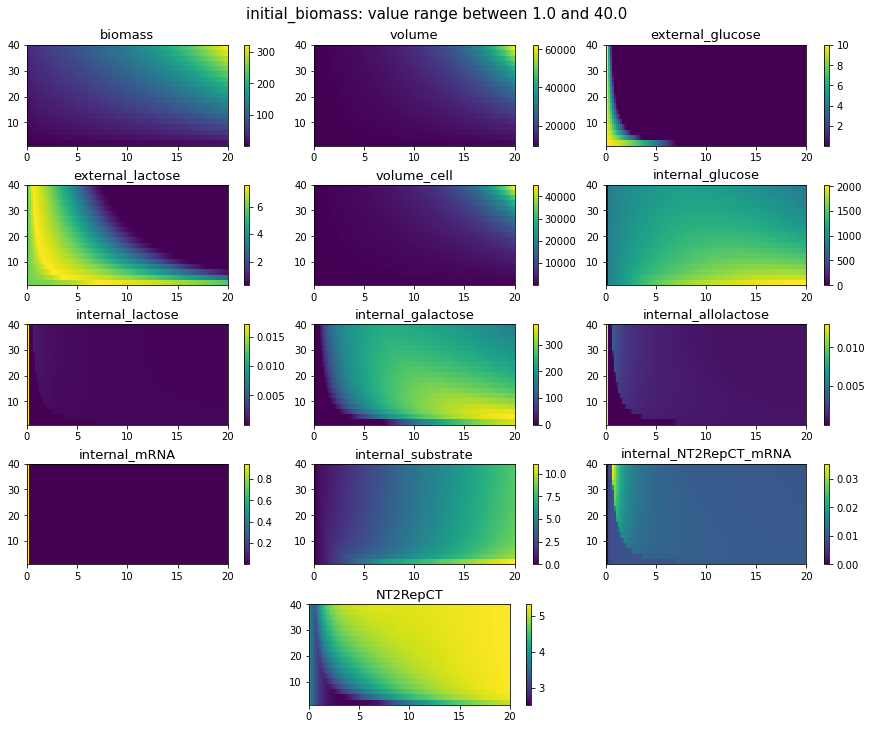
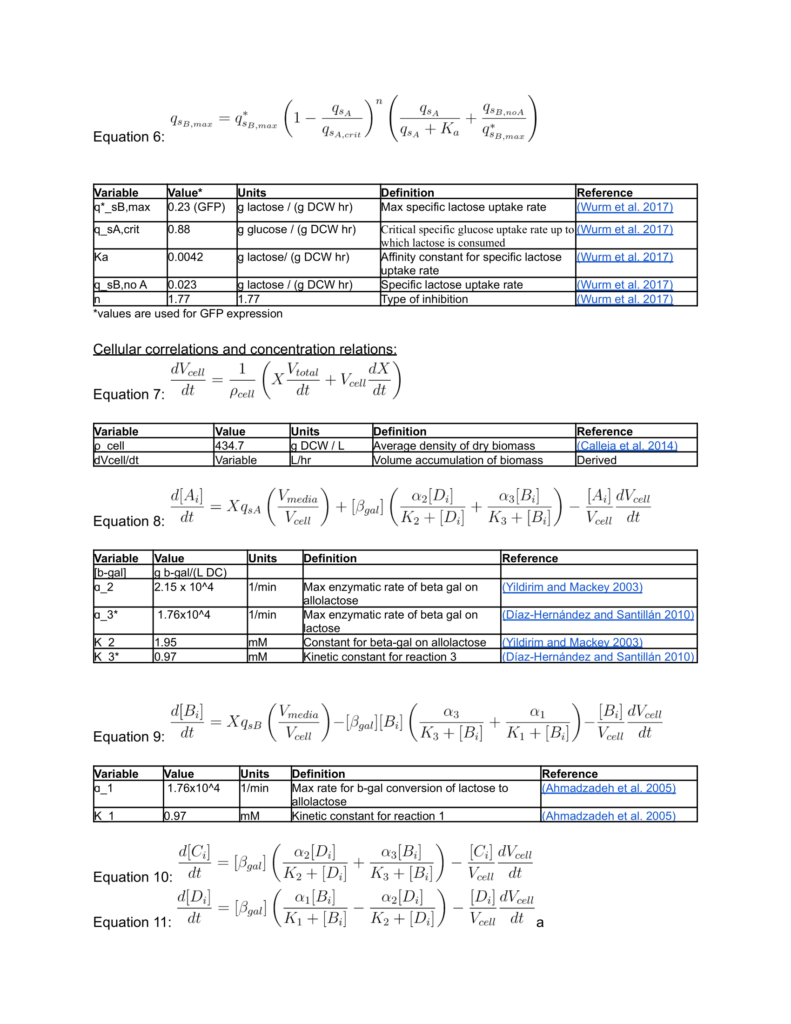
Biomass growth and Protein Synthesis Equations
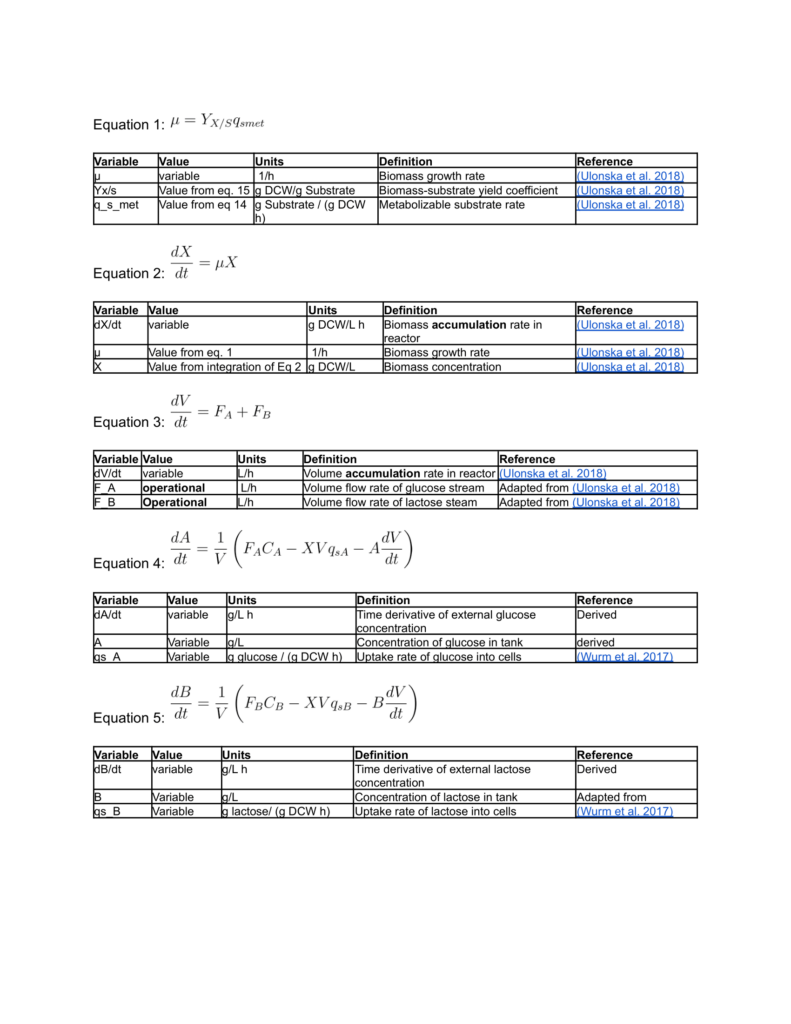


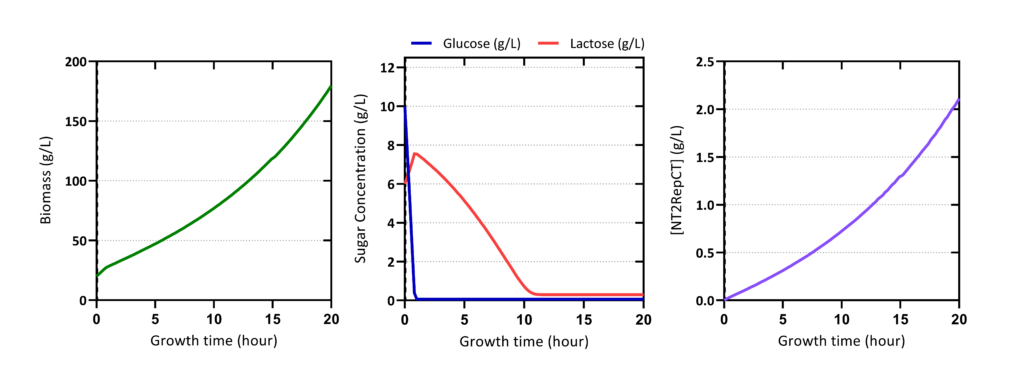

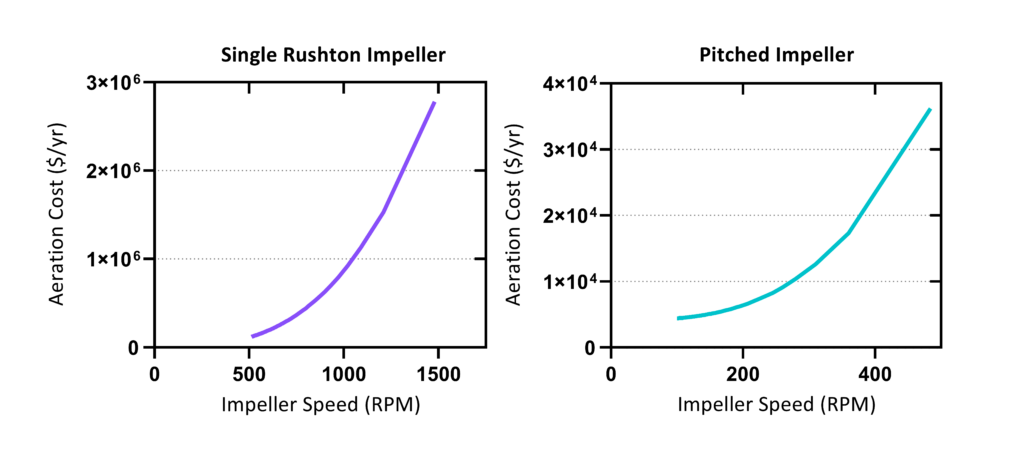
Cell Harvesting and Lysis
To perform cell lysis the designed High-Pressure Homogenizer (HPH) consists of a pump, orifice, check valve, pressure relief valve, and heat exchanger. During HPH, the fluid containing E. coli is pressurized to the desired target pressure (450 KPa). Then, the fluid is depressurized through the orifice, which causes a shear force resulting in cell lysis. The check valve is in place to prevent any backflow of the fluid, followed by a pressure relief valve which is a safety measure. Lastly, the fluid passes through a heat exchanger to account for any temperature gained during high-pressure homogenization.
Validation
To optimize the orifice and validate if the homogenizing pressure calculated is sufficient for cell lysis, the following equation from the literature was used:
The shear stress required for cell lysis for E. Coli is 1810 Pa, which was achieved from our designed HPH.
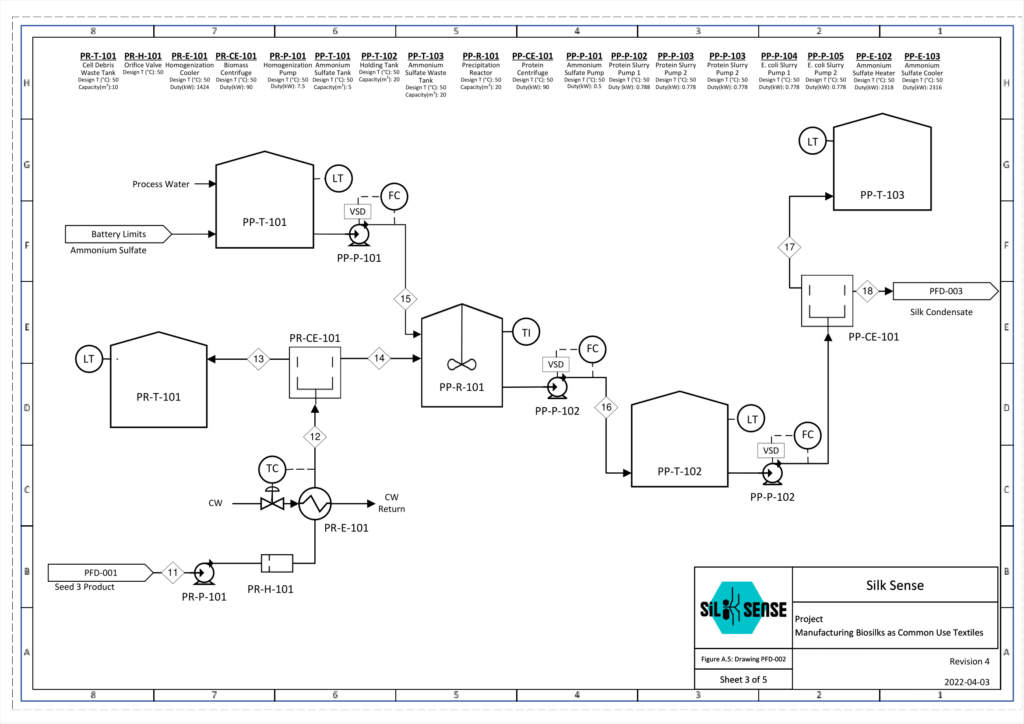
Protein Recovery and Purification
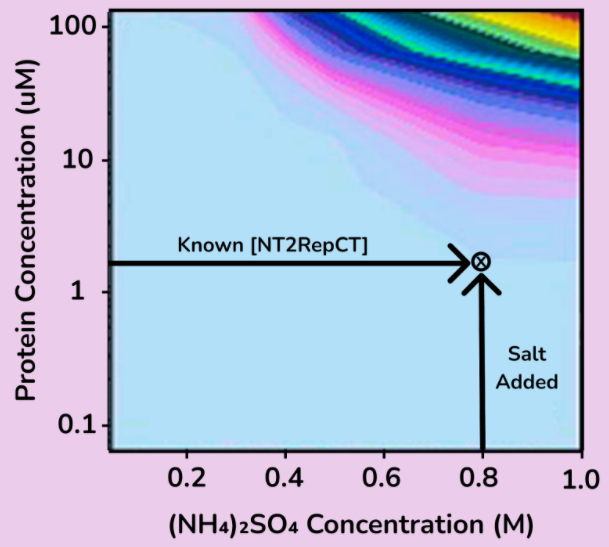
To purify our spider silk protein of interest, NT2RepCT, ammonium sulfate precipitation is used. Ammonium sulfate is a salt, and when added to a solution it can reduce the solubility of certain molecules, thus allowing them to precipitate or ‘salt out’. Salt concentrations can be specifically chosen to salt out molecules of interest, while other residual proteins stay in solution. By doing this, we can separate NT2RepCT from residual intracellular E. coli proteins, which will allow us to create a protein solution of higher purity.
Validation
Within a stirred reactor, a 3.8M ammonium sulfate solution is added to the bulk protein solution until a molarity of 0.8M ammonium sulfate is reached. A value of 0.8M was chosen based on calculations made using a kosmotrophic graph from literature. This allows NT2RepCT to precipitate and salt out, while the majority of the E. coli proteins present remain in the solution. To calculate the amount of ammonium sulfate required for this process, a mass balance was obtained from the literature and provided a value of 1,347 kg ammonium sulfate per batch.
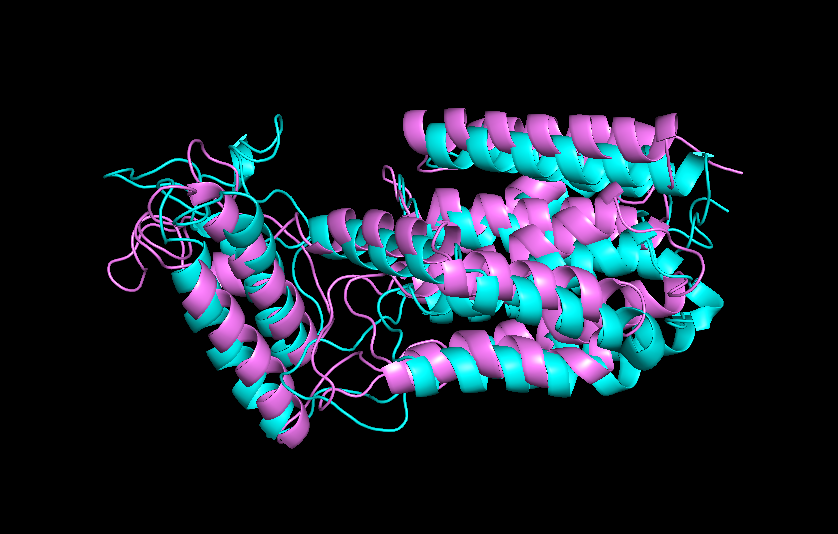
Further process design will focus on the incorporation of thermally-based recycle strategy to recovery residual silk protein and ammonium sulfate. Initial designs using computational molecular dynamics software (GROMACS) shows that NT2RepCT is thermostable up to 370K, suggesting that they can be purified from residual E.coli proteins by temperature sensitivity. Continued work will focus on completing the process calculations for this recycling process.
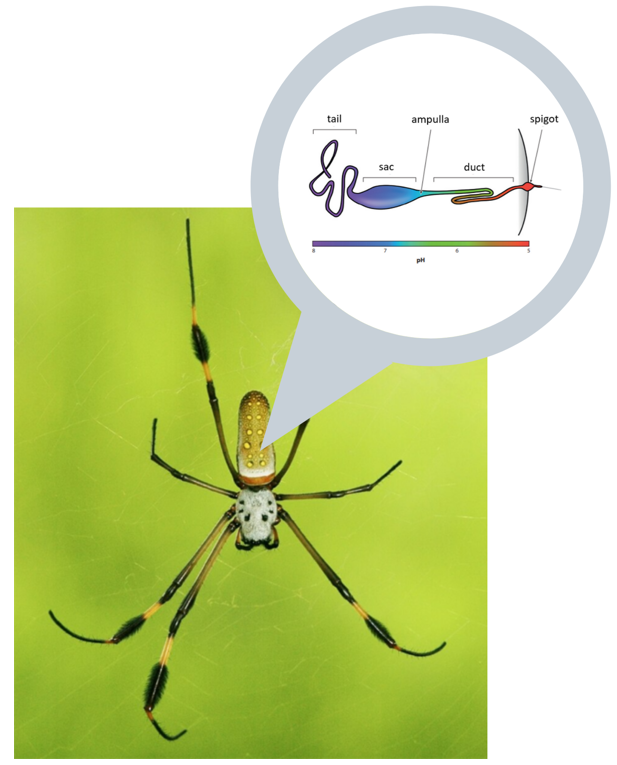
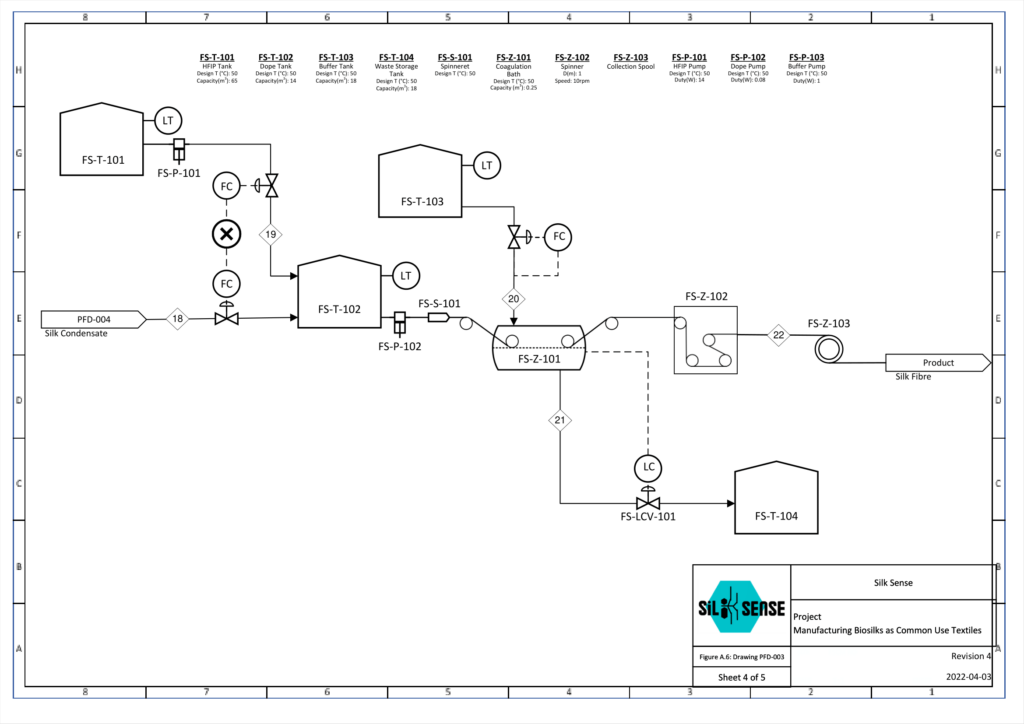
Fiber Preparation
The fiber spinning unit consists of four parts: the dope tank, the spinneret, the coagulation tank, and the collection spinner. It was designed based on a combination of nature-inspired silk formation and a wet-spinning process. This allows a high degree of alignment of proteins during spinning through a slow fiber formation. To obtain the desired mechanical properties of the silk fiber, pH, shear stress, and salt concentration are characterized. Each unit is responsible for polymerizing the protein monomers, extruding the polymer solution, enhancing the structure of the protein fiber, and collecting the product. One batch of protein which weighs up to 10kg, is spun to allow 24 batches of product per year.
Validation
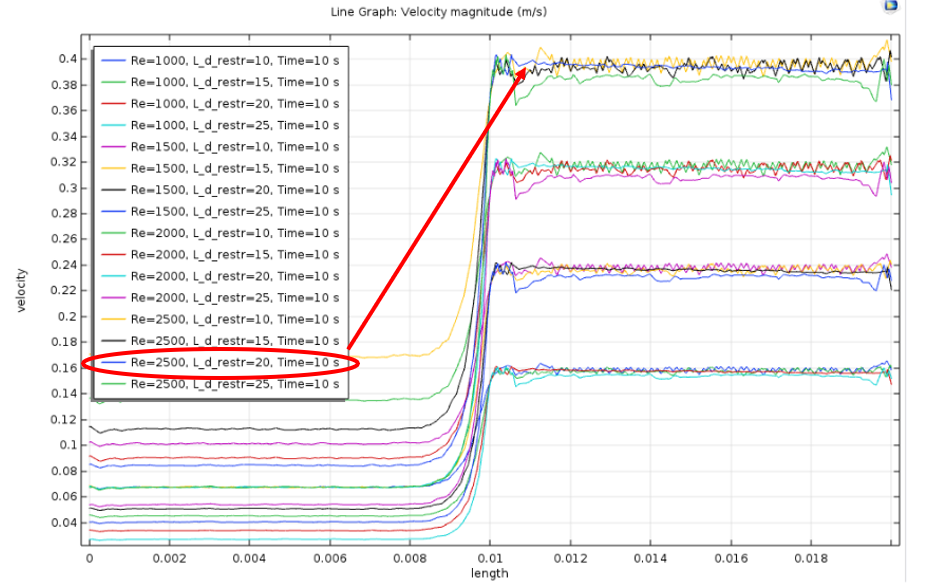
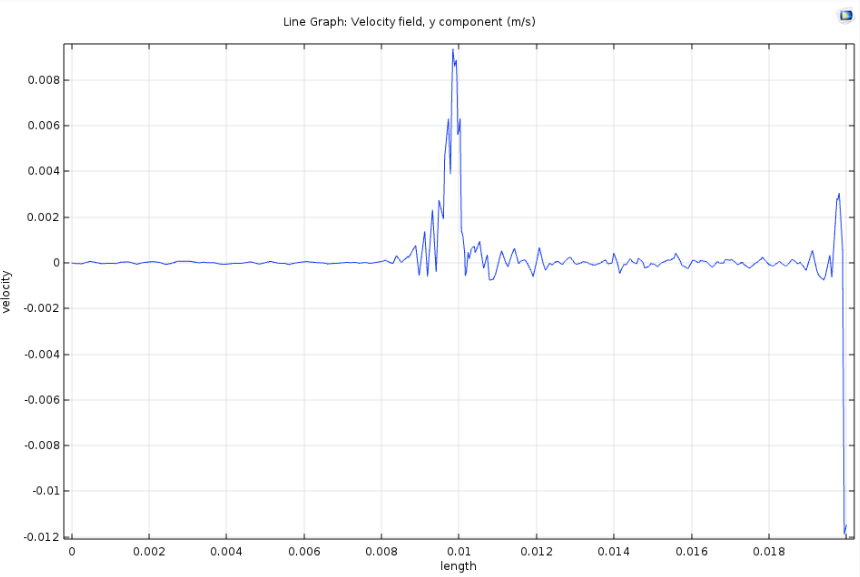
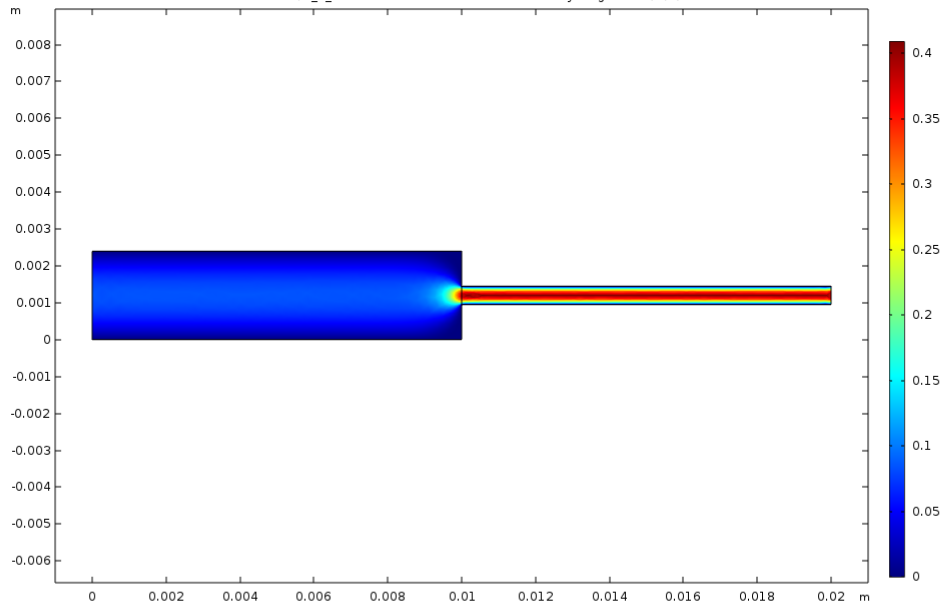
The fiber spinning unit’s production rate is dependent on the spinneret design. This component was designed and validated through fiber solidification kinetics and COMSOL software. The inlet mass flow rate and the length-to-diameter ratio were optimized to maintain the shear rate and flow regime needed to spin the spider silk fiber. This ensures that the fiber is spun continuously without rupture. The following figures demonstrate the trials conducted to evaluate the most suitable spinneret design. The spinneret design that demonstrated the least fluctuations in velocity in the x and y-coordinate was chosen. This indicates a consistent straight and laminar flow once the spinneret diameter is restricted.
Economics
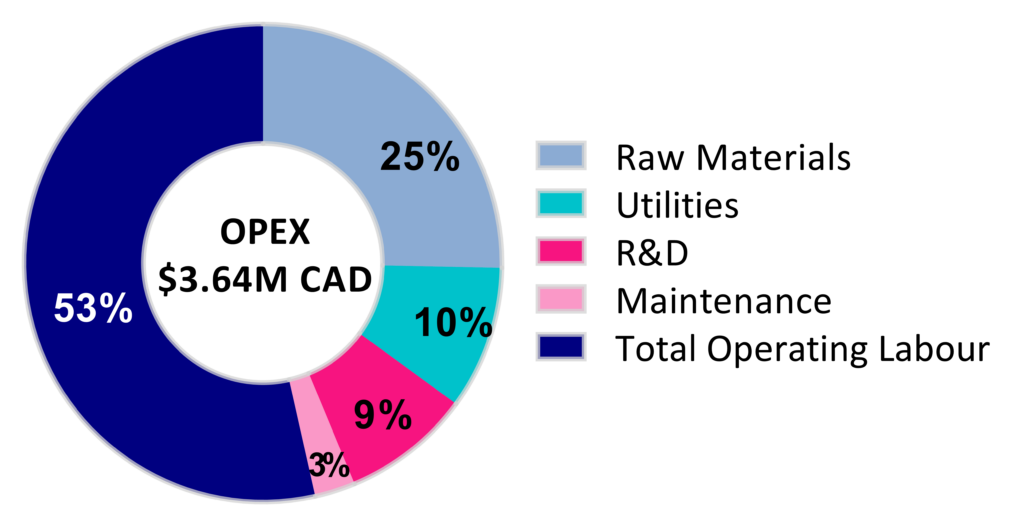
Silk Sense designed an economically feasible process with a plant lifetime of 30 years and a rate of return of 6.06%. Our plant design produces 24 batches annually and annual revenue of about $8.1M CDN. The following figure is a breakdown of our operational expenses, in which 3.64 million Canadian dollars are required to run the plant.
Sensitivity Analysis
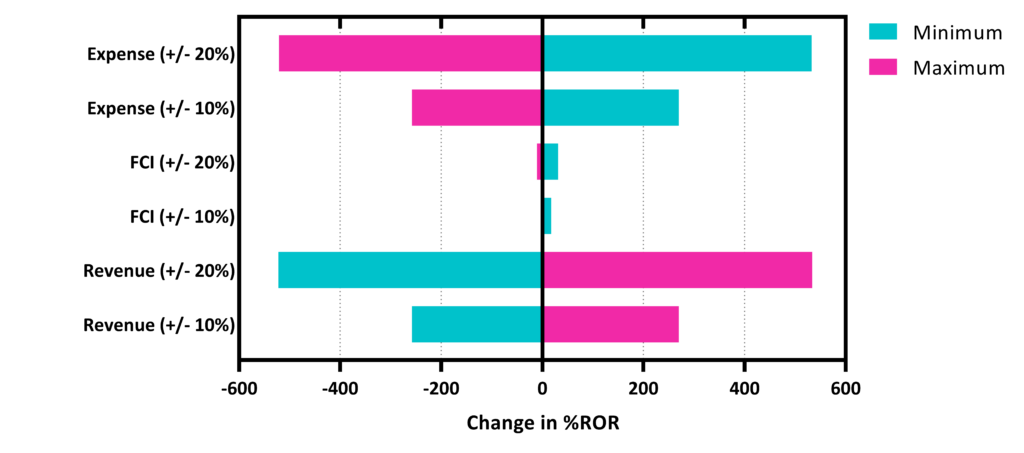
- Indicates that Revenue and Expense are the major contributors to profit margin.
- Utility and module costs were optimized to reduce expenses.
- Minimum profitable sale price can be attained by studying the changes in FCI, expense, and revenue – applied to current economics.
Social, Safety, and Environmental
Social Impacts
- Local Sustainable Products Available
- Positive Impact on Employment Rate
- Diversify the Dependency of Oil and Gas in Alberta
Operational Health and Safety
A HAZOP was conducted on the plant
- Well ventilated areas for use of toxic chemicals
- Proper use of PPE
- Responsible disposal of waste streams
- Eyewash stations and showers in close proximity
- Implementation of control loops
Environmental Impacts



Meet Our Team Members
Gabriel Oblea
Gabriel is in his final year of chemical engineering at the University of Calgary. He has experience in the upstream oil and gas industry through his internships with Imperial Oil and PureChem Services. Internship provided him opportunities in design work, optimization, and surveillance in the in situ operation site in Cold Lake. Gabriel is devoted to safety, sustainability, and finding solutions with the utmost integrity. His work in Silk Sense involves economics and the fiber spinning unit.
Kira Wasylak
Over the course of the year, Kira focused on designing the purification unit in the downstream process, and additionally, managed the process’s overall mass balance by connecting the upstream and downstream units together. Kira’s long-term career goal is to work in a leadership position within the chemical or biomedical industry. In her free time, Kira loves being active by running, hiking, skiing, and playing hockey.
Juan Sebastian Alvarez
Sebastian has dedicated his career to studying the interface of chemical engineering and synthetic biology, working towards developing solutions for environmental and biomedical challenges. He has extensive research experience through his work at iGEM Calgary, FREDsense Technologies, and the BPB Research Lab. For Silk Sense, his work has focused on managing project direction and goals, as well as the model-based design of silk-producing microorganisms.
Anna Ulrich
Anna is a final year chemical and biomedical engineering student who is passionate about improving health, energy, and environmental outcomes. She has experience in biomedical engineering research with the Lewis Research Group and Complex Fluids Lab at the University of Calgary as well as downstream oil and gas experience with Husky Energy in Lloydminster. Anna’s work with Silk Sense has focused on bioreactor equipment design and plantwide process flow drawings.
Rida Zafar
Rida is pursuing chemical engineering with a biomedical engineering specialization at the University of Calgary. Along with biomedical engineering research experience, she has experience in the midstream oil and gas industry from her internship at TC Energy. Rida’s focus is to build a career that involves the integration of sustainability into the oil and gas industry. She led the environmental, health & safety, and social impacts of Silk Sense. She also designed the cell lysis unit which consists of the high-pressure homogenizer and centrifuge.
Partners, Mentors, and Acknowledgements
We want to thank our supervisor, Dr. Jinguang Hu, for providing us with invaluable advice on biochemical processes and resources to successfully model our design. We would also like to thank our course coordinator, Dr. Hector De La Hoz Siegler, and our course instructor, Dr. Michael Foley, for guiding us in unit sizing and process flow design.
We would also like to thank Anika Zaman for her support on project and presentation design. Andrew Symes, for his support and insights into protein dynamics and molecular modelling. Rigel Tormon and Sean Ulrich for their insights and evaluation on the overall project design, as well as practice for question and answer periods.
Our Photo Gallery

References
[1] B. Resnick, “More than ever, our clothes are made of plastic. Just washing them can pollute the oceans,” Sep. 19, 2018. https://www.vox.com/the-goods/2018/9/19/17800654/clothes-plastic-pollution-polyester-washing-machine (accessed Mar. 07, 2022).
[2] S. R. Lone, V. Kumar, J. R. Seay, D. L. Englert, and H. T. Hwang, “Mass Transfer and Rheological Characteristics in a Stirred Tank Bioreactor for Cultivation of Escherichia coli BL21,” Biotechnology and Bioprocess Engineering, vol. 25, no. 5, pp. 766–776, Sep. 2020, doi: 10.1007/s12257-020-0028-3.
[3] A. Berenjian, Ed., Essentials in Fermentation Technology. Springer, Cham, 2019.
[4] – P. Mohammadi et al., “Controllable coacervation of recombinantly produced spider silk protein using kosmotropic salts,” Journal of Colloid and Interface Science, vol. 560, pp. 149–160, Feb. 2020, doi: 10.1016/j.jcis.2019.10.058.
[5] – P. Wingfield, “Protein Precipitation Using Ammonium Sulfate,” Current Protocols in Protein Science, pp. A.3F.1–A.3F.8, Sep. 1998, doi: 10.1002/0471140864.psa03fs13.
[6] Image created with BioRender.com
[7] D. J. Wurm, L. Veiter, S. Ulonska, B. Eggenreich, C. Herwig, and O. Spadiut, “The E. coli pET expression system revisited-mechanistic correlation between glucose and lactose uptake,” Appl. Microbiol. Biotechnol., vol. 100, no. 20, pp. 8721–8729, Oct. 2016.
[8] A. Ahmadzadeh, A. Halasz, V. Kumar, S. Prajna, and A. Jadbabaie, “Analysis of the Lactose metabolism in E. coli using sum-of-squares decomposition,” in Proceedings of the 44th IEEE Conference on Decision and Control, Dec. 2005, pp. 879–884.
[9] S. Ulonska, D. Waldschitz, J. Kager, and C. Herwig, “Model predictive control in comparison to elemental balance control in an E. coli fed-batch,” Chem. Eng. Sci., vol. 191, pp. 459–467, Dec. 2018.
[10] C. Mercader et al., “Kinetics of fiber solidification,” Proc. Natl. Acad. Sci. U. S. A., vol. 107, no. 43, pp. 18331–18335, Oct. 2010.
[11] T. Arndt, P. R. Laity, J. Johansson, C. Holland, and A. Rising, “Native-like Flow Properties of an Artificial Spider Silk Dope,” ACS Biomater Sci Eng, vol. 7, no. 2, pp. 462–471, Feb. 2021.







